
MTase methylates specific DNA sequences (recognition sites) in the host genome. 
Classical R-M systems of Types I–III include a DNA methyltransferase (MTase) and a restriction endonuclease (REase). 
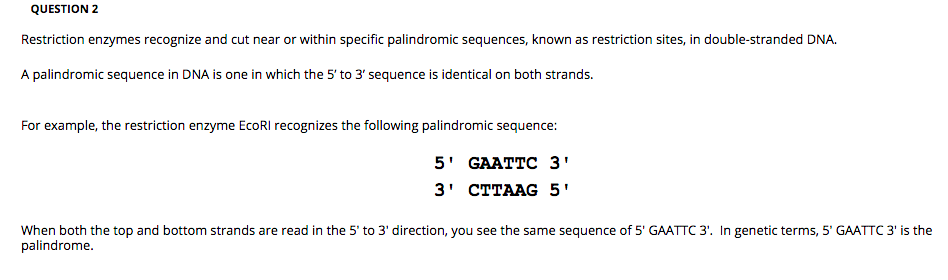
R-M systems are divided into four types (I–IV). Restriction-modification (R-M) systems were discovered and characterized as bacterial systems defending cells from an invasion of foreign DNA, e.g., phage DNA. The drastic difference between site avoidance for orthodox Type II R-M systems and R-M systems of other types can be explained by a higher rate of specificity changes or a less self-toxicity of the latter. At the same time, a significant underrepresentation of a site may be a sign of presence of the corresponding R-M system in this organism or in its ancestors for a long time. Presence of an R-M system without an underrepresentation of its site may indicate that the R-M system was acquired recently. The avoidance of orthodox Type II R-M system recognition sites in prokaryotic genomes is a widespread phenomenon. Sites of Type I, IIС/G, IIM, III, and IV R-M systems are not avoided in vast majority of cases. An analysis of closely related bacteria shows that such avoidance can be a trace of lost R-M systems. We also found cases of site avoidance in absence of the corresponding R-M systems in the genome. The sites of orthodox R-M systems that are encoded in host genomes for a long time are avoided more often (up to 100 % in certain cohorts) than the sites of recently acquired ones. We have shown that the recognition site avoidance correlates with the lifespan of R-M systems. Thus it is possible to talk about the lifespan of an R-M system in a genome. It is known that prokaryotes can acquire and lose R-M systems. Only Type II R-M systems consisting of independently acting endonuclease and methyltransferase (called ‘orthodox’ here) cause avoidance of their sites, both palindromic and asymmetric, in corresponding prokaryotic genomes the avoidance takes place for ~ 50 % of 1774 studied cases. We have studied all known recognition sites in thousands of prokaryotic genomes and found factors that influence their avoidance. However the phenomenon has not been investigated systematically for all presently available genomes and annotated R-M systems. Avoidance of palindromic recognition sites of Type II restriction-modification (R-M) systems was shown for many R-M systems in dozens of prokaryotic genomes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
